Bacteria are capable of ravaging an organism's body at a pace that medications may be unable to keep up with. Such is the virulence of these pathogens. Providing an extraordinary insight behind their ability to spread rapidly, a new study has found that bacterial groups can spread with increased rapidity on surfaces when the individuals within the group move more slowly. In other words, the slower they move, the faster they spread.
In the study that provides a better understanding of the spread of bacterial infections within the body, the researchers examined the Pseudomonas aeruginosa, a lethal species of bacteria that cause infections in both plants and animals. They found that individual bacteria that were designed to move faster were less successful than strains that were slower and moved in tightly-packed groups.
A Lethal Pathogen
Pseudomonas aeruginosa can be pathogenic in both plants and animals. In plants, they are known to cause diseases such as soft rot where sections of the plant are turned into mushy liquids by the bacteria for its easy absorption of the plant's nutrients. However, in animals and human beings, they operate as 'opportunistic' pathogens that affect individuals who have existing infections or are immunocompromised.

In human beings, they cause infections in the lungs, gastrointestinal and urinary tracts, and other regions. They also infect wounds, especially burns, and lead to several infections of the blood. Also, P. aeruginosa is a multidrug-resistant pathogen, which makes infections caused by it notoriously difficult to treat. It navigates across surfaces utilizing tiny grappling hook-like appendages known as 'pili'.
Several animals and such as birds and fishes exhibit effective movement and survival in groups. Therefore, understanding the locomotory capacity of the bacteria could offer an inward look into their ability to spread.
Slow and Steady Wins the Race
For the study, the authors compared the movement of P. aeruginosa and its 'hyperpilated' mutant individuals designed to move faster. They employed a combination of different techniques such as genetics, mathematics, and advanced tracking algorithms that were capable of monitoring the movements of several thousand cells. Through this, the scientists demonstrated that when the fast-moving engineered bacteria collided with each other, they rotated vertically and were left stuck.
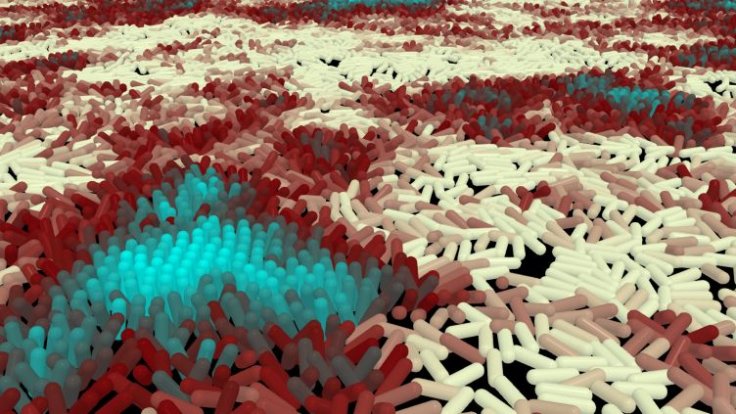

However, the slower-moving strain of P. aeruginosa remained to lie downwards when they collided with each other, thereby, facilitating them to keep moving ahead. Hence, the slow-moving bacteria can venture into new territories, obtain increased nutrients, and most importantly, win the survival race against faster cells. The findings, therefore, suggest that the evolution of the slow and restrained movement of bacteria is advantageous to the entire group, instead of individual cells.
Comparing bacterial locomotion in groups to that of human beings, Dr. William Durham, corresponding author of the study, said in a statement, "In contrast, bacteria have evolved to move carefully and effectively in crowds, likely because their neighbors tend to be genetically identical, so there is no conflict of interest. Bacteria accomplish this by moving more slowly than their top speed."
Similar to Liquid Crystals
In order to delve deeper into this bacterial phenomenon, the authors employed a theory that was developed fundamentally to learn more about materials called liquid crystals. Dr. Oliver Meacock, lead author of the study, explained, "Liquid crystals are everywhere around us, from smartphone screens to mood rings."
He added: "Although we initially didn't expect that the mathematical tools developed to understand these human-made materials could be applied to living systems, our findings show that they can also shed light on the challenges faced by microbes."









